Targeting the Biology of Aging
Aging is the biggest modifiable risk factor for most chronic diseases.1 There is now significant evidence that we can rejuvenate cells by partial epigenetic reprogramming, and thereby reverse and prevent many of these diseases. Tapping into this single, powerful aspect of biology provides us with a platform that can be applied to address a wide range of diseases.

Transformational Technology
As we age, our DNA sequence (genetic code) stays mostly the same throughout life. But the epigenetic code (which controls which genes are turned on or off) changes over time, contributing to gene expression changes that can lead to disease.
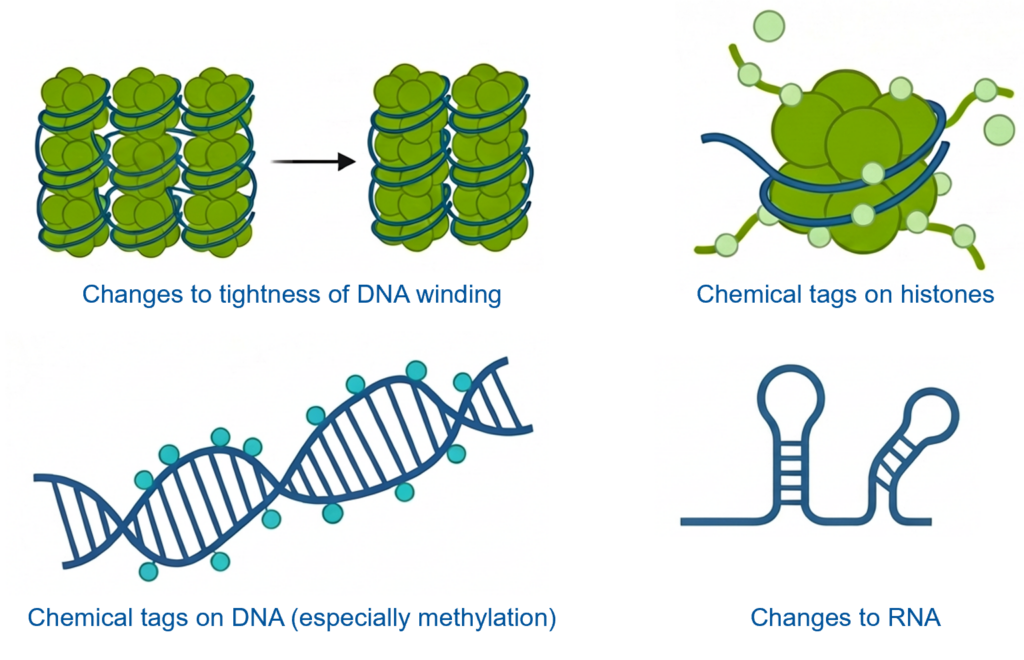
Epigenetic changes do not alter the DNA sequence itself. Instead, they affect how DNA is packaged and used by the cell. These changes include:
- How tightly DNA is wound (making genes more or less accessible)
- Chemical tags added to proteins called histones, around which DNA is wrapped (such as acetylation, methylation, ubiquitylation, sumoylation, and phosphorylation)
- Chemical tags added directly to DNA (especially DNA methylation)
- Modifications to RNA molecules that interact with DNA and influence gene activity
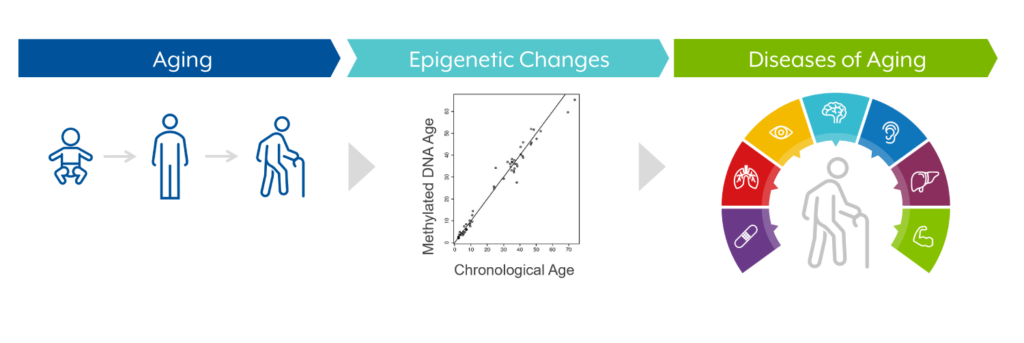
 These epigenetic changes may result from lifestyle factors (e.g., smoking, drinking, etc.), aging, disease, and injury.
These epigenetic changes may result from lifestyle factors (e.g., smoking, drinking, etc.), aging, disease, and injury.
Life Biosciences’ therapeutic approach targets these epigenetic alterations by utilizing three transcription factors – OCT4, SOX2, and KLF4 (referred to as “OSK”) – to partially reprogram the epigenome and restore aged cells to a younger, healthier state, enabling more effective cell function.
Mechanism of Action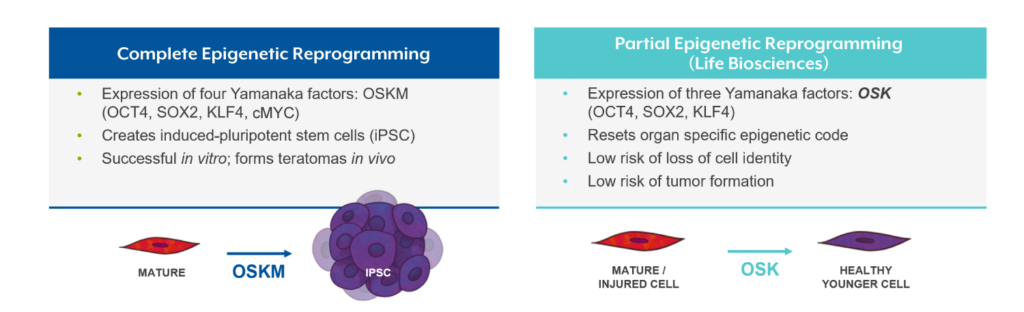
Technology That Transcends Organs
There is evidence that epigenetic changes are implicated across the vast breadth of age-related diseases, with just some of the many publications that show this connection illustrated below. There is growing evidence that reversing these epigenetic changes to restore cells to their original state could provide therapeutic benefit across these diseases. As such, Life Biosciences’ science is potentially a highly repeatable platform: while each drug candidate is a different drug product (e.g., using different vectors and promoters), the core mechanism (cellular rejuvenation using OSK) is a shared approach across diseases.
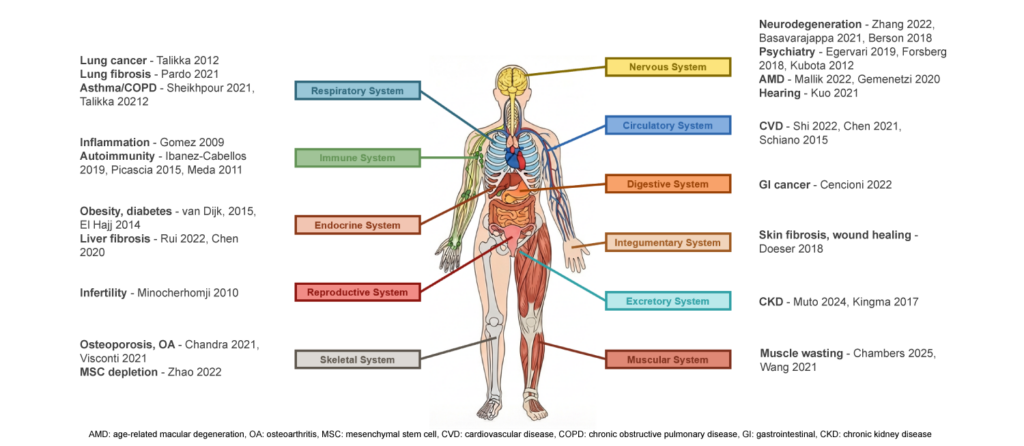
——
Berson “Epigenetic regulation in neurodegenerative diseases”, Trends Neurosci 2018
Cencioni “Gastrointestinal cancer patient nutritional management: from specific needs to novel epigenetic dietary approaches”, Nutrients 2022
Chambers “Widespread rewiring of the DNA methylome and transcriptome with decelerated methylation age after regeneration in aged skeletal muscle”, ARDD poster 2025
Chandra “Skeletal aging and osteoporosis: mechanisms and therapeutics”, Int J Mol Sci 2021
Chen “The effects of epigenetic modification on the occurrence and progression of liver diseases and the involved mechanism”, Exp Rev Gastro Hep 2020
Chen “Reversible reprogramming of cardiomyocytes to a fetal state drives heart regeneration in mice”, Science 2021
Doeser “Reduction of fibrosis and scar formation by partial reprogramming in vivo”, Stem Cells 2018
Egervari “Molecular windows into the human brain for psychiatric disorders”, Mol Psych 2019
El Hajj “Epigenetics and life-long consequences of an adverse nutritional and diabetic intrauterine environment”, Reproduction 2014
Forsberg “Epigenetics and cerebral organoids: promising directions in autism spectrum disorders”, Trans Psych 2018
Gemenetzi “Epigenetics in agerelated macular degeneration: new discoveries and future perspectives”, Cell and Mol Life Sci 2020
Gomez “Epigenetics and periodontal disease: future perspectives”, Inflamm Res 2009
Ibanez-Cabellos “Epigenetic regulation in the pathogenesis of Sjogren syndrome and rheumatoid arthritis”, Front Gen 2019
Kingma “The physiopathology of cardiorenal syndrome: a review of the potential contributions of inflammation”, J Cardio Dev Dis 2017
Kubota “Epigenetic understanding of gene-environment interactions in psychiatric disorders: a new concept of clinical genetics”, Clin Epigen 2012
Mallik “Novel epigenetic clock biomarkers of age-related macular degeneration”, Front Med 2022
Meda “The epigenetics of autoimmunity” Cell and Mol Immunol 2011
Minocherhomji “Epigenetic regulatory mechanisms associated with infertility” Obstet and Gynec Intl 2010
Muto “Epigenetic reprogramming driving successful and failed repair in acute kidney injury”, Sci Advances 2024
Pardo “The interplay of the genetic architecture, aging, and environmental factors in the pathogenesis of idiopathic pulmonary fibrosis”, Am J Resp Cell Mol Bio 2021
Picascia “Epigenetic control of autoimmune diseases: from bench to bedside”, Clin Immunol 2015
Rui “Reprogramming of hepatic metabolism and microenvironment in nonalcoholic steatohepatits”, Ann Rev Nutr 2022
Schiano “Epigenetic-related therapeutic challenges in cardiovascular disease”, Trends Pharmacol Sci 2015
Sheikhpour “A review of epigenetic changes in asthma: methylation and acetylation”, Clin Epigen 2021
Shi “Epigenetic regulation in cardiovascular disease: mechanisms and advances in clinical trials”, Signal Trans and Target Ther 2022
Talikka “Genomic impact of cigarette smoke, with application to three smoking-related diseases”, Crit Rev Toxicol 2012
van Dijk “Epigenetics and human obesity”, Intl J Obesity 2015
Visconti “DNA methylation signatures of bone metabolism in osteoporosis and osteoarthritis aging-related diseases: an updated review”, Int J Mol Sci 2021
Wang “In vivo partial reprogramming of myofibers promotes muscle regeneration by remodeling the stem cell niche”, Nat Comms 2021
Zhang “Targeting epigenetics as a promising therapeutic strategy for treatment of neurodegenerative diseases”, Biochem Pharm 2022
Zhao “Epigenetic therapy targeting bone marrow mesenchymal stem cells for age-related bone diseases”, Stem Cell Res and Therapy 2022

